PROPT Park-Ramirez bioreactor: Difference between revisions
No edit summary |
|||
| (One intermediate revision by one other user not shown) | |||
| Line 12: | Line 12: | ||
<math> J = x_1(t_f)*x_5(t_f) </math> | <math> J = x_1(t_f)*x_5(t_f) </math> | ||
subject to: | subject to: | ||
<math> \frac{dx_1}{dt} = g_1*(x_2-x_1)-\frac{u}{x_5}*x_1 ,</math> | <math> \frac{dx_1}{dt} = g_1*(x_2-x_1)-\frac{u}{x_5}*x_1 ,</math> | ||
<math> \frac{dx_2}{dt} = g_2*x_3-\frac{u}{x_5}*x_2 ,</math> | <math> \frac{dx_2}{dt} = g_2*x_3-\frac{u}{x_5}*x_2 ,</math> | ||
<math> \frac{dx_3}{dt} = g_3*x_3-\frac{u}{x_5}*x_3 ,</math> | <math> \frac{dx_3}{dt} = g_3*x_3-\frac{u}{x_5}*x_3 ,</math> | ||
<math> \frac{dx_4}{dt} = -7.3*g_3*x_3+\frac{u}{x_5}*(20-x_4) ,</math> | <math> \frac{dx_4}{dt} = -7.3*g_3*x_3+\frac{u}{x_5}*(20-x_4) ,</math> | ||
<math> \frac{dx_5}{dt} = u ,</math> | <math> \frac{dx_5}{dt} = u ,</math> | ||
with: | with: | ||
<math> g_1 = 4.75*\frac{g_3}{(0.12+g_3)} ,</math> | <math> g_1 = 4.75*\frac{g_3}{(0.12+g_3)} ,</math> | ||
<math> g_2 = \frac{x_4}{(0.1+x4)}*exp(-5*x_4) ,</math> | <math> g_2 = \frac{x_4}{(0.1+x4)}*exp(-5*x_4) ,</math> | ||
<math> g_3 = 21.87*\frac{x_4}{(x_4+0.4)*(x_4+62.5)} ,</math> | <math> g_3 = 21.87*\frac{x_4}{(x_4+0.4)*(x_4+62.5)} ,</math> | ||
where x1 and x2 are, respectively, the concentration of the secreted protein and the total protein (l-1), x3 is the culture cell density (g l-1), x4 is the substrate concentration (g l-1), x5 is the holdup volume (l), u is the nutrient (glucose) feed rate (l h-1), and J is the mass of protein produced (in arbitrary units). The initial conditions are: | where x1 and x2 are, respectively, the concentration of the secreted protein and the total protein (l-1), x3 is the culture cell density (g l-1), x4 is the substrate concentration (g l-1), x5 is the holdup volume (l), u is the nutrient (glucose) feed rate (l h-1), and J is the mass of protein produced (in arbitrary units). The initial conditions are: | ||
<math> x(t_0) = [0 \ 0 \ 1 \ 5 \ 1]' </math> | <math> x(t_0) = [0 \ 0 \ 1 \ 5 \ 1]' </math> | ||
For final time t_f = 15 h, and the following constraints on the control variable: | For final time t_f = 15 h, and the following constraints on the control variable: | ||
<math> 0 <= u <= 2 </math> | <math> 0 <= u <= 2 </math> | ||
<source lang="matlab"> | <source lang="matlab"> | ||
| Line 231: | Line 242: | ||
[[File:parkBioReactor_01.png]] | [[File:parkBioReactor_01.png]] | ||
[[Category:PROPT Examples]] | |||
Latest revision as of 05:32, 14 February 2012
|
This page is part of the PROPT Manual. See PROPT Manual. |
Dynamic optimization of chemical and biochemical processes using restricted second-order information 2001, Eva Balsa-Canto, Julio R. Banga, Antonio A. Alonso Vassilios S. Vassiliadis
Case Study I: Park-Ramirez bioreactor
Problem description
This case study deals with the optimal production of secreted protein in a fed-batch reactor. It was originally formulated by Park and Ramirez (Park & Ramirez, 1988) and it has also been considered by other researchers (Vassiliadis, 1993; Yang & Wu, 1994; Banga, Irizarry & Seider, 1995; Luus, 1995; Tholudur & Ramirez, 1997). The objective is to maximize the secreted heterologous protein by a yeast strain in a fed-batch culture. The dynamic model accounts for host-cell growth, gene expression, and the secretion of expressed polypeptides. The mathematical statement is as follows:
Find u(t) over t in [t0,t_f] to maximize:
subject to:
with:
where x1 and x2 are, respectively, the concentration of the secreted protein and the total protein (l-1), x3 is the culture cell density (g l-1), x4 is the substrate concentration (g l-1), x5 is the holdup volume (l), u is the nutrient (glucose) feed rate (l h-1), and J is the mass of protein produced (in arbitrary units). The initial conditions are:
For final time t_f = 15 h, and the following constraints on the control variable:
% Copyright (c) 2007-2008 by Tomlab Optimization Inc.Problem setup
toms tSolve the problem, using a successively larger number collocation points
for n=[20 40 80 120]
p = tomPhase('p', t, 0, 15, n);
setPhase(p);
tomStates z1 z2 z3 z4 z5
tomControls u
if n>20
% Interpolate an initial guess for the n collocation points
x0 = {icollocate({z1 == z1opt; z2 == z2opt
z3 == z3opt; z4 == z4opt; z5 == z5opt})
collocate(u == uopt)};
else
x0 = {};
end
% Box constraints
cbox = {icollocate({z1 <= 3
z2 <= 3; 0 <= z3 <= 4
0 <= z4 <= 10; 0.5 <= z5 <= 25})
0 <= collocate(u) <= 2.5};
% Boundary constraints
cbnd = initial({z1 == 0; z2 == 0; z3 == 1
z4 == 5; z5 == 1});
% Various constants and expressions
g3 = 21.87*z4./(z4+.4)./(z4+62.5);
g1 = 4.75*g3./(0.12+g3);
g2 = z4./(0.1+z4).*exp(-5*z4);
% ODEs and path constraints
ceq = collocate({
dot(z1) == g1.*(z2-z1)-u./z5.*z1
dot(z2) == g2.*z3-u./z5.*z2
dot(z3) == g3.*z3-u./z5.*z3
dot(z4) == -7.3*g3.*z3+u./z5.*(20-z4)
dot(z5) == u});
% Secreted protein must be less than total protein
% proteinlimit = {z1 <= z2};
% Objective
if n == 120
objective = -final(z1)*final(z5)+var(diff(collocate(u)));
else
objective = -final(z1)*final(z5);
endSolve the problem
options = struct;
options.name = 'Park Bio Reactor';
solution = ezsolve(objective, {cbox, cbnd, ceq}, x0, options);
% Optimal z and u, to use as starting guess in the
% next iteration
z1opt = subs(z1, solution);
z2opt = subs(z2, solution);
z3opt = subs(z3, solution);
z4opt = subs(z4, solution);
z5opt = subs(z5, solution);
uopt = subs(u, solution);Problem type appears to be: qpcon
Time for symbolic processing: 0.48856 seconds
Starting numeric solver
===== * * * =================================================================== * * *
TOMLAB - TOMLAB Development license 999007. Valid to 2011-12-31
=====================================================================================
Problem: --- 1: Park Bio Reactor f_k -31.891604942090204000
sum(|constr|) 0.000000000222835644
f(x_k) + sum(|constr|) -31.891604941867367000
f(x_0) 0.000000000000000000
Solver: snopt. EXIT=0. INFORM=1.
SNOPT 7.2-5 NLP code
Optimality conditions satisfied
FuncEv 1 ConstrEv 102 ConJacEv 102 Iter 61 MinorIter 911
CPU time: 0.171601 sec. Elapsed time: 0.172000 sec.
Problem type appears to be: qpcon
Time for symbolic processing: 0.48207 seconds
Starting numeric solver
===== * * * =================================================================== * * *
TOMLAB - TOMLAB Development license 999007. Valid to 2011-12-31
=====================================================================================
Problem: --- 1: Park Bio Reactor f_k -29.523702508277267000
sum(|constr|) 0.000000000531678452
f(x_k) + sum(|constr|) -29.523702507745590000
f(x_0) -31.891604942089948000
Solver: snopt. EXIT=0. INFORM=1.
SNOPT 7.2-5 NLP code
Optimality conditions satisfied
FuncEv 1 ConstrEv 75 ConJacEv 75 Iter 58 MinorIter 588
CPU time: 0.436803 sec. Elapsed time: 0.434000 sec.
Problem type appears to be: qpcon
Time for symbolic processing: 0.4837 seconds
Starting numeric solver
===== * * * =================================================================== * * *
TOMLAB - TOMLAB Development license 999007. Valid to 2011-12-31
=====================================================================================
Problem: --- 1: Park Bio Reactor f_k -31.866762161406442000
sum(|constr|) 0.000000000726937524
f(x_k) + sum(|constr|) -31.866762160679503000
f(x_0) -29.523702508277417000
Solver: snopt. EXIT=0. INFORM=1.
SNOPT 7.2-5 NLP code
Optimality conditions satisfied
FuncEv 1 ConstrEv 137 ConJacEv 137 Iter 120 MinorIter 784
CPU time: 4.586429 sec. Elapsed time: 4.584000 sec.
Problem type appears to be: qpcon
Time for symbolic processing: 0.51719 seconds
Starting numeric solver
===== * * * =================================================================== * * *
TOMLAB - TOMLAB Development license 999007. Valid to 2011-12-31
=====================================================================================
Problem: --- 1: Park Bio Reactor f_k -32.616144466384739000
sum(|constr|) 0.000000000312031161
f(x_k) + sum(|constr|) -32.616144466072711000
f(x_0) -31.477810607457531000
Solver: snopt. EXIT=0. INFORM=1.
SNOPT 7.2-5 NLP code
Optimality conditions satisfied
FuncEv 1 ConstrEv 134 ConJacEv 134 Iter 116 MinorIter 930
CPU time: 46.815900 sec. Elapsed time: 16.326000 sec.
end
t = subs(collocate(t),solution);
z1 = collocate(z1opt);
z2 = collocate(z2opt);
z3 = collocate(z3opt);
z4 = collocate(z4opt);
z5 = collocate(z5opt);
u = collocate(uopt);Plot result
subplot(2,1,1)
plot(t,z1,'*-',t,z2,'*-',t,z3,'*-',t,z4,'*-',t,z5,'*-');
legend('z1','z2','z3','z4','z5');
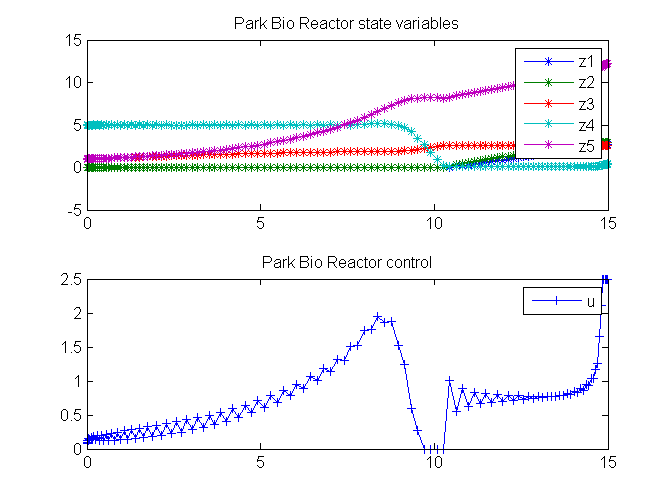
title('Park Bio Reactor state variables');
subplot(2,1,2)
plot(t,u,'+-');
legend('u');
title('Park Bio Reactor control');








![{\displaystyle x(t_{0})=[0\ 0\ 1\ 5\ 1]'}](https://wikimedia.org/api/rest_v1/media/math/render/svg/c9fbcb4f5193a58243fc1c61f61ddbca4a794a50)