PROPT Lee-Ramirez Bioreactor: Difference between revisions
(Created page with "{{Part Of Manual|title=the PROPT Manual|link=PROPT Manual}} Dynamic optimization of chemical and biochemical processes using restricted second-order information 200...") |
No edit summary |
||
| Line 1: | Line 1: | ||
{{Part Of Manual|title=the PROPT | {{Part Of Manual|title=the PROPT Manual|link=[[PROPT|PROPT Manual]]}} | ||
Dynamic optimization of chemical and biochemical processes using restricted second-order information 2001, Eva Balsa-Canto, Julio R. Banga, Antonio A. Alonso Vassilios S. Vassiliadis | Dynamic optimization of chemical and biochemical processes using restricted second-order information 2001, Eva Balsa-Canto, Julio R. Banga, Antonio A. Alonso Vassilios S. Vassiliadis | ||
| Line 269: | Line 269: | ||
</pre> | </pre> | ||
[[File:leeBioReactor_01.png]] | |||
[[File:leeBioReactor_02.png]] | |||
Revision as of 14:24, 2 November 2011
|
This page is part of the PROPT Manual. See PROPT Manual. |
Dynamic optimization of chemical and biochemical processes using restricted second-order information 2001, Eva Balsa-Canto, Julio R. Banga, Antonio A. Alonso Vassilios S. Vassiliadis
Case Study II: Lee-Ramirez bioreactor
Problem description
This problem considers the optimal control of a fed-batch reactor for induced foreign protein production by recombinant bacteria, as presented by Lee and Ramirez (1994) and considered afterwards by Tholudur and Ramirez (1997) and Carrasco and Banga (1998). The objective is to maximize the profitability of the process using the nutrient and the inducer feeding rates as the control variables. Three different values for the ratio of the cost of inducer to the value of the protein production (Q) were considered.
The mathematical formulation, following the modified parameter function set presented by Tholudur and Ramirez (1997) to increase the sensitivity to the controls, is as follows:
Find u1(t) and u2(t) over t in [t0 t_f] to maximize:
subject to:
where the state variables are the reactor volume (x1), the cell density (x2), the nutrient concentration (x3), the foreign protein concentration (x4), the inducer concentration (x5), the inducer shock factor on cell growth rate (x6) and the inducer recovery factor on cell growth rate (x7). The two control variables are the glucose rate (u1) and the inducer feeding rate (u2). Q is the ratio of the cost of inducer to the value of the protein production, and the final time is considered fixed as t_f = 10 h. The model parameters were described by Lee and Ramirez (1994). The initial conditions are:
The following constraints on the control variables are considered:
% Copyright (c) 2007-2008 by Tomlab Optimization Inc.Problem setup
toms t
for n=[20 35 55 85]
p = tomPhase('p', t, 0, 10, n);
setPhase(p);
tomStates z1 z2 z3s z4 z5 z6 z7
% Declaring u as "states" makes it possible to work with their
% derivatives.
tomStates u1 u2
% Scale z3 by 40
z3 = z3s*40;
% Initial guess
if n == 20
x0 = {icollocate({z1 == 1; z2 == 0.1
z3 == 40; z4 == 0; z5 == 0
z6 == 1; z7 == 0})
icollocate({u1==t/10; u2==t/10})};
else
x0 = {icollocate({z1 == z1opt
z2 == z2opt; z3 == z3opt
z4 == z4opt; z5 == z5opt
z6 == z6opt; z7 == z7opt})
icollocate({u1 == u1opt
u2 == u2opt})};
end
% Box constraints
cbox = {mcollocate({0 <= z1; 0 <= z2
0 <= z3; 0 <= z4; 0 <= z5})
0 <= collocate(u1) <= 1
0 <= collocate(u2) <= 1};
% Boundary constraints
cbnd = initial({z1 == 1; z2 == 0.1
z3 == 40; z4 == 0; z5 == 0
z6 == 1; z7 == 0});
% Various constants and expressions
c1 = 100; c2 = 0.51; c3 = 4.0;
Q = 0;
t1 = 14.35+z3+((z3).^2/111.5);
t2 = 0.22+z5;
t3 = z6+0.22./t2.*z7;
g1 = z3./t1.*(z6+z7*0.22./t2);
g2 = 0.233*z3./t1.*((0.0005+z5)./(0.022+z5));
g3 = 0.09*z5./(0.034+z5);
% ODEs and path constraints
ceq = collocate({
dot(z1) == u1+u2
dot(z2) == g1.*z2-(u1+u2).*z2./z1
dot(z3) == u1./z1.*c1-(u1+u2).*z3./z1-g1.*z2/c2
dot(z4) == g2.*z2-(u1+u2).*z4./z1
dot(z5) == u2*c3./z1-(u1+u2).*z5./z1
dot(z6) == -g3.*z6
dot(z7) == g3.*(1-z7)});
% Objective
J = -final(z1)*final(z4)+Q*integrate(u2);
spenalty = 0.1/n; % penalty term to yield a smoother u.
objective = J + spenalty*integrate(dot(u1)^2+dot(u2)^2);Solve the problem
options = struct;
options.name = 'Lee Bio Reactor';
solution = ezsolve(objective, {cbox, cbnd, ceq}, x0, options);
% Optimal z, u for starting point
z1opt = subs(z1, solution);
z2opt = subs(z2, solution);
z3opt = subs(z3, solution);
z4opt = subs(z4, solution);
z5opt = subs(z5, solution);
z6opt = subs(z6, solution);
z7opt = subs(z7, solution);
u1opt = subs(u1, solution);
u2opt = subs(u2, solution);Problem type appears to be: qpcon
Time for symbolic processing: 0.85283 seconds
Starting numeric solver
===== * * * =================================================================== * * *
TOMLAB - TOMLAB Development license 999007. Valid to 2011-12-31
=====================================================================================
Problem: --- 1: Lee Bio Reactor f_k -6.158551034261496900
sum(|constr|) 0.000000279053783607
f(x_k) + sum(|constr|) -6.158550755207713200
f(x_0) 0.001000000000000196
Solver: snopt. EXIT=0. INFORM=1.
SNOPT 7.2-5 NLP code
Optimality conditions satisfied
FuncEv 1 ConstrEv 744 ConJacEv 744 Iter 573 MinorIter 3217
CPU time: 2.542816 sec. Elapsed time: 2.538000 sec.
Problem type appears to be: qpcon
Time for symbolic processing: 0.86391 seconds
Starting numeric solver
===== * * * =================================================================== * * *
TOMLAB - TOMLAB Development license 999007. Valid to 2011-12-31
=====================================================================================
Problem: --- 1: Lee Bio Reactor f_k -6.149281027809802700
sum(|constr|) 0.000002151758382440
f(x_k) + sum(|constr|) -6.149278876051420500
f(x_0) -6.161215441652444700
Solver: snopt. EXIT=0. INFORM=1.
SNOPT 7.2-5 NLP code
Optimality conditions satisfied
FuncEv 1 ConstrEv 548 ConJacEv 548 Iter 486 MinorIter 1855
CPU time: 5.709637 sec. Elapsed time: 5.969000 sec.
Problem type appears to be: qpcon
Time for symbolic processing: 0.84929 seconds
Starting numeric solver
===== * * * =================================================================== * * *
TOMLAB - TOMLAB Development license 999007. Valid to 2011-12-31
=====================================================================================
Problem: --- 1: Lee Bio Reactor f_k -6.148479046961865500
sum(|constr|) 0.000000948376662244
f(x_k) + sum(|constr|) -6.148478098585203000
f(x_0) -6.151342657272863300
Solver: snopt. EXIT=0. INFORM=1.
SNOPT 7.2-5 NLP code
Optimality conditions satisfied
FuncEv 1 ConstrEv 508 ConJacEv 508 Iter 443 MinorIter 2428
CPU time: 16.520506 sec. Elapsed time: 17.194000 sec.
Problem type appears to be: qpcon
Time for symbolic processing: 0.83028 seconds
Starting numeric solver
===== * * * =================================================================== * * *
TOMLAB - TOMLAB Development license 999007. Valid to 2011-12-31
=====================================================================================
Problem: --- 1: Lee Bio Reactor f_k -6.149330296208792600
sum(|constr|) 0.000000385398306198
f(x_k) + sum(|constr|) -6.149329910810486400
f(x_0) -6.149452089871609000
Solver: snopt. EXIT=0. INFORM=1.
SNOPT 7.2-5 NLP code
Optimality conditions satisfied
FuncEv 1 ConstrEv 689 ConJacEv 689 Iter 616 MinorIter 2438
CPU time: 83.694536 sec. Elapsed time: 85.082000 sec.
end
t = subs(collocate(t),solution);
z1 = subs(collocate(z1),solution);
z2 = subs(collocate(z2),solution);
z3 = subs(collocate(z3),solution);
z4 = subs(collocate(z4),solution);
z5 = subs(collocate(z5),solution);
z6 = subs(collocate(z6),solution);
z7 = subs(collocate(z7),solution);
u1 = subs(collocate(u1),solution);
u2 = subs(collocate(u2),solution);Plot result
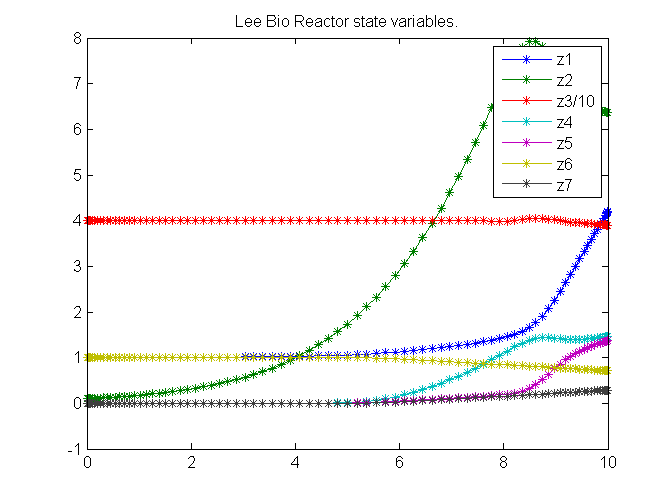
figure(1)
plot(t,z1,'*-',t,z2,'*-',t,z3/10,'*-',t,z4,'*-' ...
,t,z5,'*-',t,z6,'*-',t,z7,'*-');
legend('z1','z2','z3/10','z4','z5','z6','z7');
title('Lee Bio Reactor state variables.');
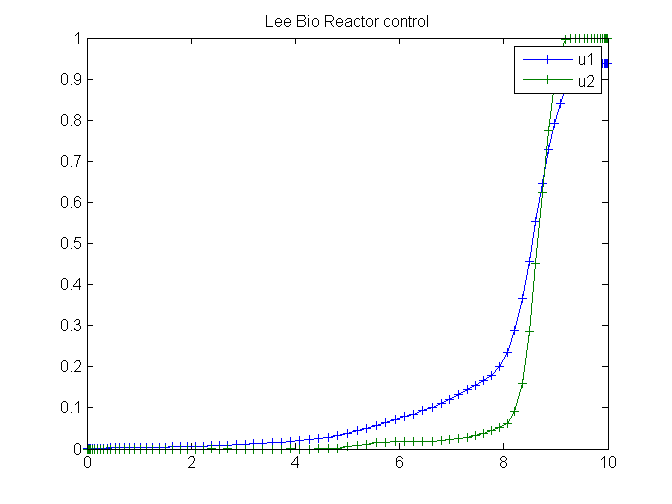
figure(2)
plot(t,u1,'+-',t,u2,'+-');
legend('u1','u2');
title('Lee Bio Reactor control');
disp('J = ');
disp(subs(J,solution));J = -6.1510

















![{\displaystyle x(t_{0})=[1\ 0.1\ 40\ 0\ 0\ 1\ 0]'}](https://wikimedia.org/api/rest_v1/media/math/render/svg/f6b688a78a64862acef297acac2cd449298fbc5b)